视神经损伤
-
Figure 1|Heterogeneity of ocular macrophages in health and disease.
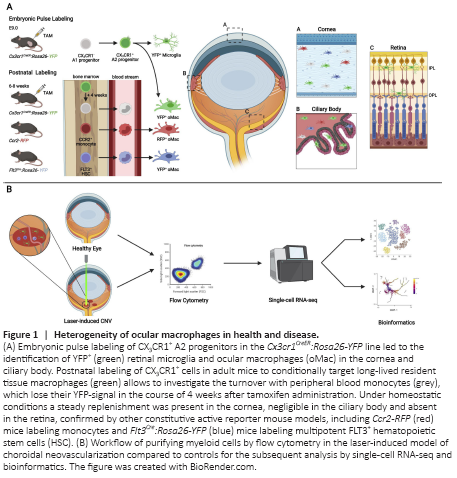
In two separate studies, our group and others used multiple fate mapping systems to provide evidence for a prenatal origin of myeloid cells in the eye. By applying the Runx1MER-Cre-MER:Rosa26-YFP model, O’Koren et al. (2019) demonstrated that the administration of 4-hydroxytamoxifen at E7.5 leads to a similar amount of YFP+ cells in the retina and the brain at 8 weeks postpartum, indicating a yolk sac-derived origin as in the brain. These findings were supported by us using the Cx3cr1CreER:Rosa26-YFP mouse model, which was shown to consistently label bMG in an embryonic fate mapping approach (Goldmann et al., 2016). Taking advantage of this model in the context of the eye, our results clearly point towards a prenatal origin of retinal microglia (rMG) (Wieghofer et al., 2021). In addition, we extended our analysis towards myeloid cell populations in the ciliary body and the cornea. Here, we were able to show for the first time that cells targeted by embryonic pulse labeling were present in the ciliary body and cornea as well (Wieghofer et al., 2021). To address the question whether circulating monocytes from the definitive hematopoiesis contribute to the myeloid cell pool in different eye compartments, we used a fate mapping approach in adult mice by injecting tamoxifen into Cx3cr1CreER:Rosa26-YFP mice at the age of 6–8 weeks. Retinal microglia showed a very low turnover that is reminiscent of the longevity of bMG or CAMs (Goldmann et al., 2016, Wieghofer et al., 2021). Interestingly, we found that ciliary body macrophages also showed relatively stable YFP-labeling indicating the longevity of these cells whereas the percentage of YFP+ cells in the cornea significantly dropped over the course of time (Wieghofer et al., 2021), which is comparable to the findings in choroidal macrophages (O’Koren et al., 2019). These results were further supported by the use of Ccr2-RFP mice, where short-lived monocytes are specifically labeled, and Flt3Cre:Rosa26-YFP to target multipotent hematopoietic progenitor cells of the fetal and adult hematopoiesis. In both models, the highest recombination rates were present in the cornea and to a smaller extent in the ciliary body, whereas the recombination in the retina was negligible (Wieghofer et al., 2021). In summary, these studies redefined our understanding of the ontogeny of oMacs in different compartments of the eye and highlighted the different kinetics by which these macrophages are replenished by circulating monocytes in vivo (Figure 1A).
To characterize the myeloid cell heterogeneity in multiple eye compartments including the retina, ciliary body and cornea, we used scRNA-seq to decipher the myeloid cell landscape at single-cell resolution (Figure 1B) (Wieghofer et al., 2021). We identified several cell clusters corresponding to different myeloid cell types like microglia or other macrophages and found a significant enrichment of these clusters in different compartments of the eye (Wieghofer et al., 2021). Microglial cell clusters exhibited the homeostatic gene expression signature including Tmem119 and P2ry12 (Wieghofer et al., 2021), thereby confirming the previous results from O’Koren and colleagues (O’Koren et al., 2019). Furthermore, we were able to show that bMG and rMG exhibit strong transcriptional similarities (Wieghofer et al., 2021), which may be explained by their shared ontogeny and environment as these factors are decisive for macrophage identity. Contrarily, cornea macrophages clearly separated from microglia clusters with strong similarities to bone marrow-derived monocytes reflecting the aforementioned differences in postnatal turnover (Wieghofer et al., 2021). To investigate the role of distinct myeloid subsets in retinal pathology, we used scRNA-seq in a laser-induced model of choroidal neovascularization (CNV), a commonly used mouse model for neovascular age-related macular degeneration (Wieghofer et al., 2021). Here, we identified multiple disease-associated cell clusters being enriched at different time points after laser treatment and therefore reflecting successive changes in transcriptional profiles at different disease stages. Disease-associated microglia (DAM) were the dominant myeloid cell type in CNV lesions, as shown in CNV-induced Cx3cr1CreERT2:Rosa26-tdTomato mice, and characterized by the lower abundance of homeostatic genes and the concomitant upregulation of glycolytic enzymes like Gapdh which may indicate an altered microglial metabolism (Wieghofer et al., 2021). To get deeper insights into the spatiotemporal kinetics underlying these transcriptional changes, a trajectory analysis was performed. Interestingly, cells at day 7 after CNV-induction were more similar to the homeostatic signature than cells at day 3 which may be explained by the fact that day 3 may represent the more acute phase of CNV. In another study, single-cell profiling of microglia in a model of photoreceptor degeneration revealed the presence of DAM with the downregulation of homeostatic genes and the upregulation of Lgals3, Spp1 and Lpl (O’Koren et al., 2019). Interestingly, immunofluorescence revealed a significant enrichment of these cell clusters in the subretinal space, which is devoid of any immune cells under physiological conditions (O’Koren et al., 2019; Wieghofer et al., 2021). Of note, previous studies in the brain highlighted the importance of DAM clusters as potential targets for myeloid cell-based interventions in neurodegenerative and neuroinflammatory diseases (Ajami et al., 2018; Jord?o et al., 2019; Masuda et al., 2019). As bulk and scRNA-seq showed that rMG and bMG are transcriptionally very similar, it is tempting to hypothesize that the DAM clusters identified in ocular diseases may overlap with DAMs in the brain (O’Koren et al., 2019; Wieghofer et al., 2021). Thus, this approach may help us to identify molecular signatures, that are conserved across diseases in the brain and the eye and may therefore facilitate the development of drugs intended to reverse the pathological alterations in myeloid cells.