中国神经再生研究(英文版) ›› 2026, Vol. 21 ›› Issue (8): 3807-3814.doi: 10.4103/NRR.NRR-D-25-00107
携带特定线粒体单倍群帕金森病患者星形胶质细胞的缺氧及TTR基因表达失调:单核细胞转录组解析
Hypoxia and TTR dysregulation in astrocytes from Parkinson’s disease with a specific mitochondrial haplogroup: A single-cell analysis
Junhao Wang1, 2, #, Wenxuan Du1, 2, #, Xinyi Chen1, 2, Hao Wu1, 2, Ganqiang Liu1, 2, 3, *
- 1Shenzhen Key Laboratory of Systems Medicine in Inflammatory Diseases, School of Medicine, Shenzhen Campus of Sun Yat-sen University, Shenzhen, Guangdong Province, China;
2Neurobiology Research Center, School of Medicine, Shenzhen Campus of Sun Yat-sen University, Shenzhen, Guangdong Province, China;
3Guangdong Provincial Key Laboratory of Brain Function and Disease, Guangzhou, Guangdong Province, China
摘要:
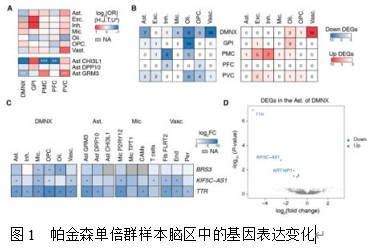
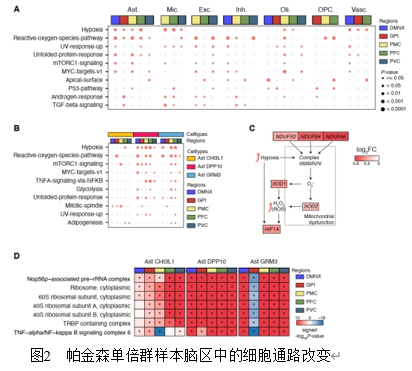
线粒体DNA(mtDNA)变异与帕金森病的认知进展相关,但mtDNA变异或单倍型群如何参与这一过程的机制尚不明确。此研究通过通过对不同线粒体单倍群帕金森病患者死后不同脑区的单核细胞转录组解析发现,线粒体单倍群H亚群新皮层区域中星形胶质细胞(Ast)亚型Ast CHI3L1和Ast GRM3的比例发生显著改变。值得注意的是,H亚群X神经背侧运动核(DMNX)区域中TTR基因表达显著下调。通路分析显示星形胶质细胞中存在异常的缺氧和活性氧(ROS)环境,而蛋白质复合物分析表明Ast亚型中核糖体亚基复合物的一致性显著升高。通过构建加权有向全转录组基因调控网络,发现H亚群星形胶质细胞亚型中SP1和HOXA5的活性发生显著变化。此外,观察到多个转录因子对TTR基因转录调控的广泛失调。此外,H 型单倍群的帕金森病患者在某些脑区显示出网络功能连接性增强。这项数据驱动的研究强调了线粒体 DNA 单倍群通过细胞组成变化、差异基因表达、通路紊乱和基因调控网络参与帕金森病认知进展的潜在机制。此研究构建mtDNA单倍群H驱动帕金森病认知障碍的多维度调控框架,为帕金森病精准诊疗提供了新思路。
https://orcid.org/0000-0002-1921-9542 (Ganqiang Liu)